Wild Relatives of Sweetpotato
Crop wild relatives represent a diverse gene pool which can contribute beneficial traits such as resistance to biotic (insects and diseases) and abiotic (drought, frost, salinity) factors to crop improvement programs. The genetic erosion and loss of habitat of these valuable crop wild relatives is occurring as human populations grow and expand and variable climate changes natural habitats too rapidly for the plants to adapt. This put pressure on us to preserve theses wild relatives in genebanks before they are permanently lost. The genebank from the International Potato Center (CIP) maintains 1,092 wild sweetpotato accessions corresponding to 67 species from 19 countries. The wild collections are conserved as populations as seed in -20°C cold chambers. This includes the Batatas Series which consists of 13 species.
Seed regeneration of wild sweetpotato is undertaken using a few fundamental principles to produce enough seed for distribution while still maintaining as much allelic diversity as possible from the original collection. During regeneration, the accession is also characterized using standard genebank descriptors. Flower formation of sweetpotato, Ipomoea batatas (L.) Lam., and its wild relatives is challenging because flowering is dependent on various environmental conditions, some of which are still active areas of research to determine what is needed for flowering and seed formation. Flowering can vary widely in different seasons/years from poor to abundant. Some accessions produce little or no flowers like those from the species I. ramossisima, I. tiliaceae, and I. tabascana. For species and accessions which produce few flowers, the induction of flowering is often successful by an assortment of methods including a shortened photoperiod, increased humidity, chemicals, grafting onto other Ipomoea species or increased nutrient fertilization.
Seed Regeneration of Wild Relatives of Sweetpotato






Ploidy Determination of Sweetpotato Wild Relatives by Flow Cytometry
Flow cytometry is a technique used for the determination of ploidy and nuclear DNA content. A major advantage of flow cytometry is the large number of nuclei analyzed in a very short period of time. This allows a rapid screening of large amounts of samples from breeding experiments, in-vitro regeneration, routine analysis for identity or compatibility checking or characterization studies in a genebank. Chromosome counting is performed for results validation.

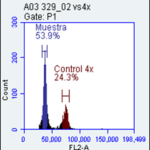
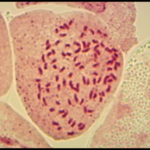
Taxonomic classification of Batata series and ploidy level(s)
| Ipomoea spp. | Ploidy 2n (x=15) |
|---|---|
| I. batatas | 6x, 4x |
| I. cordatotriloba | 2x, 4x |
| I. cynanchifolia | 2x |
| I. x grandifolia | 2x |
| I. lacunosa | 2x |
| I. x leucantha | 2x |
| I. littoralis | 2x |
| I. ramosissima | 2x |
| I. tabascana | 4x |
| I. tenuissima | 2x |
| I. tiliacea | 4x |
| I. trifida | 2x, 4x, 6x |
| I. triloba | 2x |
Contact
Genoveva Rossel
Sweetpotato Curator
g.rossel@cgiar.org


